Loading component...
Depression is a dark horse.
The disease often goes unnoticed, but affects work performance, social interaction and the ability to take pleasure in everyday life. According to the National Center for Biotechnology Information, antidepressants only help around 50 percent of those who struggle with depression and anxiety and, even when they are effective, scientists have yet to understand how they work in the brain.

But groundbreaking research in the lab of Michigan State University scientist A.J. Robison, associate professor in the Department of Physiology and MSU’s Neuroscience Program, is directing some new rays of light onto the molecular, cellular and circuit-level mechanisms underlying depression-like diseases.
The results were recently published in Nature Communications.
“In this paper, we perform the first ever CRISPR-based gene editing [a genetic engineering technique in molecular biology by which the genomes of living organisms may be modified] in a single circuit between two areas of the mouse brain,” explained Robison about the culmination of five years of research funded by the National Institutes of Mental Health. “We can reach into the mouse brain and manipulate specific genes in a circuit involved in depression and anxiety-like behaviors — a critical advance on the road to genetic medicine for psychiatric diseases.”
Scientists estimate there are roughly 80-100 billion neurons connecting regions of the brain. To accomplish the feat of locating and manipulating a single gene in a single circuit required new and sophisticated technology. With the expertise of co-author Rachael Neve, director of the Gene Transfer Core at Massachusetts General Hospital, they developed it.
“The key advance is that we designed a dual-vector system to manipulate a specific gene in the connections between two brain areas, and that has never been done before,” Robison said.
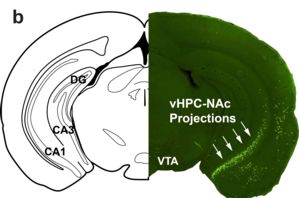
The neurons that Robison and his team zeroed in on originate in the ventral hippocampus (vHPC), a deep-seated structure that projects to regions in the brain important in stress susceptibility, mood and social avoidance. Neurons rooted in the vHPC reach out with branch-like structures called axons to connect with the nucleus accumbens, or NAc. The completed circuit is regulated by the star of the pioneering paper, the transcription factor known as DFosB.
Using the viral vector technology specifically designed and packaged by Neve, the team split the CRISPR system in half. Half of the system, inert on its own, was an enzyme that can mutate DNA in the vHPC. The other half, a guide RNA, was sent to all cells that project to the NAc and tells the enzyme where to bind and the specific gene to mutate. Only those cells specific to the circuit from the vHPC to the NAc got both halves, triggering the enzyme to bind with and turn off a single gene: FosB.
“When the FosB gene was turned off in the neurons, we were able to get a circuit-specific behavioral effect relevant to a disease like depression,” said Robison about the landmark discovery. “When we put it back, or rescued it within the circuit, the effect was erased.”

“One of the most exciting findings from our investigations was the circuit-specific role of the ΔFosB protein in conferring resilience to stress,” Eagle said. “We also discovered that ΔFosB altered the excitability of hippocampal circuit neurons and may be affecting long-term downstream changes that lead to changes in the activity of this circuit.” But removing DFosB permanently altered the expression of a suite of genes, in effect removing the conductor from the orchestra. To that end, the paper goes on to report in-depth experiments on DFosB largely done by the members of the Robison Lab including co-first authors Claire Manning, a 2019 neuroscience graduate, now a postdoc at Stanford University; and Andrew Eagle, a former postdoctoral researcher, now an assistant professor in the MSU Department of Physiology.

Based on the findings in the paper, the Robison Lab will continue to develop highly collaborative and cutting-edge techniques, accelerated by MSU’s newly completed Interdisciplinary Science and Technology Building. “This work is important because it elucidates a potential mechanism, namely ΔFosB, for how stress may contribute to depression,” Eagle continued. “Future clinical work may find ways to directly manipulate ΔFosB, or more likely one of its gene targets, to provide resilience to stress and decrease the incidence of depression in vulnerable people.”
“The end of this paper, which shows us measuring the changes of expression in hundreds of genes when we remove DFosB, is only the beginning of years of work for our lab,” Robison said. “Which genes are important and what are they doing in the brain? This is the challenge of a lifetime for me and my lab.”
This article is repurposed content originally featured on the College of Natural Sciences website.